| Descriptions |
pDRAW32 is a "plasmid drawing software". Following features are described on the software website
- pDRAW32 lets you enter a DNA name and coordinates for genetic elements, such as genes, to be plotted on your DNA plots
- pDRAW32 lets you "clone" fragments of DNA generated by virtual digestion with restriction enzymes and optionally blunted at one or both ends. Up to 3 fragments may be cloned at a time (can you replicate that in the lab?). Each fragment may be inverted relative to its original orientation. Genetic elements contained in the cloned fragments are transferred to the cloned DNA.
- pDRAW32 enables you to edit the sequence in various ways. You can invert the sequence or rotate it, which may generate more pleasing plots without actually altering the DNA sequence.
- Upon opening an existing sequence, entering a new sequence or generating a newly cloned sequence pDRAW32 automatically analyses it for
restriction enzyme sites
dam and dcm methylation sites
open reading frames using standard or non-standard genetic codes
oligonucleotide priming sites
... more to come
- pDRAW32 lets you select which restriction enzymes to plot in the graphical output and list in the textual output. To be selected an enzyme must fit all of the criteria's "Min/Max", "Seq type", and "Subset" OR the criteria "Explicit select". You may also chose to only select the explicitly selected enzymes.
Enzymes may be selected on the basis of how many times they cut in up to 5 different regions of the DNA (of which one is the entire sequence). If an enzyme cuts more than a given "Display cut off" limit, the hits and fragments are not listed in the textual output.
- pDRAW32 can display and print various different textual outputs
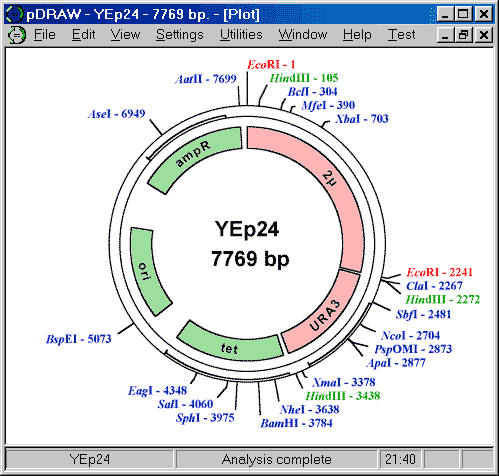
- pDRAW32 displays, prints, and exports graphical plots. The plot is based on the analysis result, the restriction enzymes and oligonucleotides you have selected, the annotation data you have entered, and the color and font settings you have entered. Circular DNA may be plotted either circular or linear. The plots contain DNA name and size, open reading frames, genetic elements you may have entered, recognition sites for restriction enzymes you have selected, and oligonucleotide priming sites. For each restriction site the name of the enzyme is plotted and optionally the position, the recognition sequence, the number of sites, and whether the particular site is inhibited by dam or dcm methylation. On the monitor the graphics may be zoomed by up to 400%.
- pDRAW32 can generate virtual agarose gel plots, plotting the generated fragment sizes generated with the selected enzymes and enzyme mixtures. Different agarose gel-percentage, run length (Min bp), and DNA molecular weight markers can be selected.
- pDRAW32 can generate a XY plot of homology between a given sequence and itself or another sequence
- pDRAW32 allows you to calculate the optimal PCR annealing temperature based on the sequence of a pair of primers. The TM calculator by default calculates TM based on the nearest neighbour method (entropy and enthalpy) but can also calculate TM by the simpler GC content or 2+4 algorithms. A number of parameters can be set, including template concentration, product length, GC content of template, and the concentrations of primer, NaCl, formamide, glycerol and DMSO.
- pDRAW32 supplies a scientific helpfile which can help you quickly lookup certain scientific facts. Most notably, you can lookup the activities of restriction enzymes in the various buffer systems from Promega, New England Biolabs, Boehringer, and Gibco/Life Technologies.
 |
- runs on different platforms
- free
|
|
|